Proteomics
"This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison."
Introduction
What is proteomics?
|
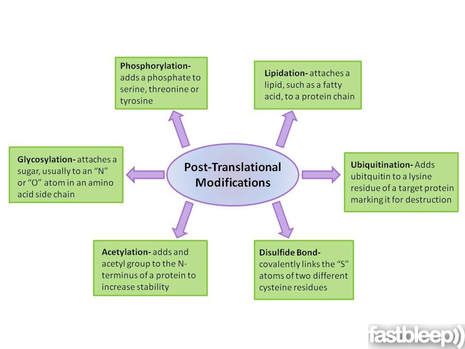
Broadly speaking, proteomics is how proteins function inside of a cell (1). Often times, proteins are influenced by post-translational modifications. Post-translational modifications include phosphorylation, methylation, acetylation, and more (2). These protein modification increase the diversity of the proteome, and are important in protein interactions and cellular signaling (2). Interestingly, the human genome contains about 20,000-25,000 genes, but thanks to post-translational modifications, over 1 million proteins (2). Post-translational modifications can be vital for proper protein function.
|
Results
Mouse snca Post-translational modifications
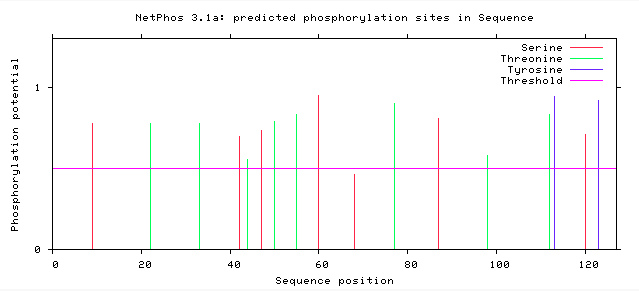
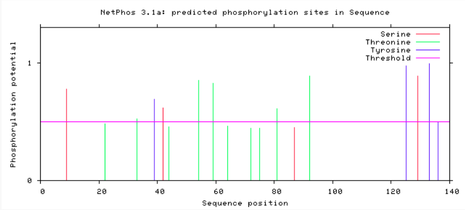
NetPhos generates predicted serine, threonine, and tyrosine within an amino acid sequence. The human SNCA and zebrafish SNCB proteins have several predicted phosphorylation sites, many of which could affect protein activity if altered. However, interestingly, there seems to be little conservation of these sites between the two homologous proteins.
Discussion
Phosphorylation can increase or turn off protein or enzyme activity. There are several predicted SNCA and SNCB phosphorylation sites, therefore creating several possible critical sites within the protein. If even one amino acid in one of these critical areas is changed or altered, proper protein activity may be compromised. Conserved sites among two different proteins may play a vital role in protein activity. These changes can cause disease, but an added challenge is that they may be difficult to detect.
References
1. What is Proteomics?-News Medical Life Sciences. (April, 2016). Retrieved from https://www.news-medical.net/life-sciences/What-is-Proteomics.aspx
2. Overview of Post-Translational Modifications (PMTs)-Thermo Fisher Scientific. Retrieved from https://www.thermofisher.com/us/en/home/life-science/protein-biology/protein-biology-learning-center/protein-biology-resource-library/pierce-protein-methods/overview-post-translational-modification.html
Predicted phosphorylation sites: NetPhos 3.1 Server. Retrieved from http://www.cbs.dtu.dk/services/NetPhos/
2. Overview of Post-Translational Modifications (PMTs)-Thermo Fisher Scientific. Retrieved from https://www.thermofisher.com/us/en/home/life-science/protein-biology/protein-biology-learning-center/protein-biology-resource-library/pierce-protein-methods/overview-post-translational-modification.html
Predicted phosphorylation sites: NetPhos 3.1 Server. Retrieved from http://www.cbs.dtu.dk/services/NetPhos/